Standard Test Statistics for OLS Models in R
with tags normality-test t-test F-test hausman-test -Model testing belongs to the main tasks of any econometric analysis. This post gives an overview of tests, which should be applied to OLS regressions, and illustrates how to calculate them in R. The focus of the post is rather on the calcuation of the tests. For a treatment of mathematical details, please, consult a standard textbook. This list of statistical tests is necessarily incomplete. However, if you have a strong opinion that a specific test is missing, feel free to leave a comment below.
Data and model
To illustrate the calculation of test statistics in R, let’s use the wage1 dataset from the wooldridge package and estimate a basic Mincer earnings function. This standard specification of earnings models explains the natural log of average hourly earnings lwage by years of education educ and experience exper. The standard specification also includes squared values of experience expersq to take into account potential decreasing marginal effects.
# Load dataset
library(wooldridge)
data("wage1")
# Estimate a model
model <- lm(lwage ~ educ + exper + expersq, data = wage1)Significance of variables
t test
t tests are used to assess the statistical significance of single variables. In R t values for each variable in a regression model are usually calculated by the summary function:
summary(model)##
## Call:
## lm(formula = lwage ~ educ + exper + expersq, data = wage1)
##
## Residuals:
## Min 1Q Median 3Q Max
## -1.96387 -0.29375 -0.04009 0.29497 1.30216
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 0.1279975 0.1059323 1.208 0.227
## educ 0.0903658 0.0074680 12.100 < 2e-16 ***
## exper 0.0410089 0.0051965 7.892 1.77e-14 ***
## expersq -0.0007136 0.0001158 -6.164 1.42e-09 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 0.4459 on 522 degrees of freedom
## Multiple R-squared: 0.3003, Adjusted R-squared: 0.2963
## F-statistic: 74.67 on 3 and 522 DF, p-value: < 2.2e-16The t values for our benchmark model indicate that, except for the constant, all variables are statistically significant. You might think about dropping the intercept term at this point, but let’s forget about this for the moment.
F test
F tests can be used to check, whether one or multiple variables in a model are statistically significant. Basically, the test compares two models with each other, where one model is a special case of the other. This means that we compare a model with more variables - the so-called unrestricted model - to a model with less but otherwise the same variables, i.e. the restriced or nested model. If the additional predictive power of the unrestricted model is sufficiently higher, the variables are jointly significant.
In our example we add the tenure variable and its square tenuresq to the model equation. This model is now the unrestricted model, which we have to estimate before we can calculate the F test.
# Estimate unrestricted model
model_unres <- lm(lwage ~ educ + exper + expersq + tenure + tenursq, data = wage1)
summary(model_unres)##
## Call:
## lm(formula = lwage ~ educ + exper + expersq + tenure + tenursq,
## data = wage1)
##
## Residuals:
## Min 1Q Median 3Q Max
## -1.96984 -0.25313 -0.03204 0.27141 1.28302
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 0.2015715 0.1014697 1.987 0.0475 *
## educ 0.0845258 0.0071614 11.803 < 2e-16 ***
## exper 0.0293010 0.0052885 5.540 4.80e-08 ***
## expersq -0.0005918 0.0001141 -5.189 3.04e-07 ***
## tenure 0.0371222 0.0072432 5.125 4.20e-07 ***
## tenursq -0.0006156 0.0002495 -2.468 0.0139 *
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 0.425 on 520 degrees of freedom
## Multiple R-squared: 0.3669, Adjusted R-squared: 0.3608
## F-statistic: 60.26 on 5 and 520 DF, p-value: < 2.2e-16The t values of the new model indicate that all variabes - including the constant - are individually, statistically significant. Thus, the variable tenure provides additional explanatory power to the model. Since tenure also enters in its squared form, we are interested in the joint significance of the tenure terms. This can be checked with an F test, which can be calculated by using the anova function in R. We only have to provide the two estimated models as arguments:
anova(model, model_unres)## Analysis of Variance Table
##
## Model 1: lwage ~ educ + exper + expersq
## Model 2: lwage ~ educ + exper + expersq + tenure + tenursq
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 522 103.790
## 2 520 93.911 2 9.8791 27.351 5.079e-12 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1As you can see from the output, the tenure terms are joinly significant as indicated by the low value in column Pr(>F).
Note that both the \(t\) test and the \(F\) test are only valid in situations, where the relation between the dependent and independent variables are linear. For non-linear cases the likelihood ratio (LR) test, the Wald (W) test, and the Lagrange multiplier (LM) test can be used.
Normality of the errors
One key assumption of many test statistics is that the errors of a model are normally distributed. If this is not the case, some of the obtained results might become questionable. There is a series of normality tests, which are also listed on Wikipedia: Normality tests. A selection of these tests is presented in the following.
Note that Kennedy (2014) mentions these tests in the context of outlier detection. This is intuitive, since outliers usually lie outside of the assumed (normal) distribution.
Tests
- Eyeball test: A rather intuitive way to test for normality is to simply look at the histogram of the model residuals:
hist(model$residuals, breaks = 20)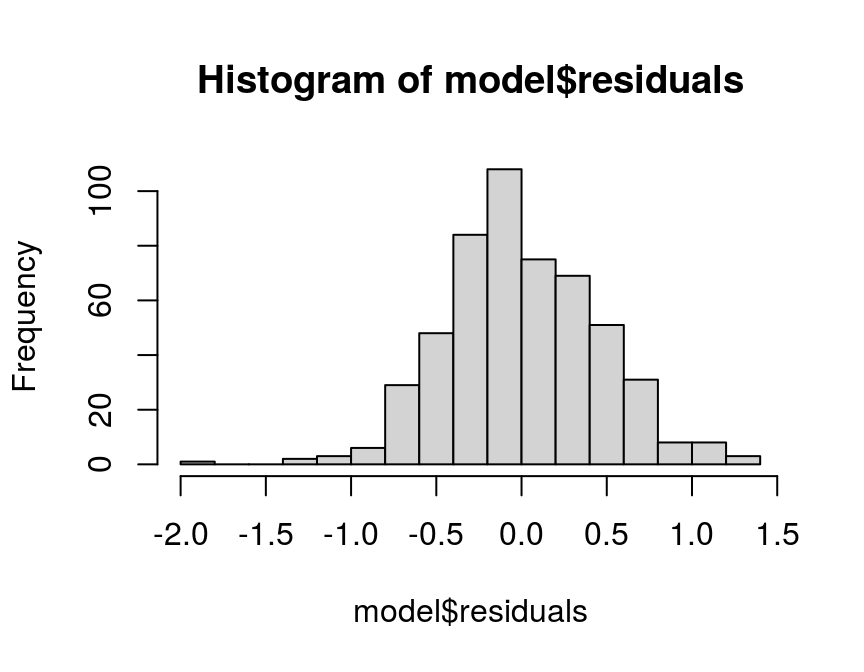
Since the shape of the histogram resembles the famous bell shape of a normal distribution, this looks pretty normal to me. However, there are more formal ways to test for normality, which are required by a more sophisticated audience than me…
- The Shapiro-Wilk test is considered to be a very reliable test for normality. It is implemented in the
shapiro.testfunction of thestatspackage:
shapiro.test(model$residuals)##
## Shapiro-Wilk normality test
##
## data: model$residuals
## W = 0.99258, p-value = 0.01031The low p-value indicates that the null hypothesis of a normal distribution is not rejected at the 1 percent level.
- The Anderson–Darling test is also a popular normality test. It also has the advantage that it can be applied to other distributions as well. To test for normal distributions in R, the
ad.testfunction of thenortestpackage can be used:
library(nortest)
ad.test(model$residuals)##
## Anderson-Darling normality test
##
## data: model$residuals
## A = 0.86991, p-value = 0.02559The low p-value also indicates that the null hypothesis of a normal distribution is not rejected at the 2.56 percent level.
- The Jarque-Bera test tests whether the skewness and kurtosis of a model’s residuals match the expected values for normal distribution. In R the function
jarque.bera.testof thetseriespackages and the functionjarque.testin themomentspackage are, among others, implementations of this test:
library(tseries)
jarque.bera.test(model$residuals)##
## Jarque Bera Test
##
## data: model$residuals
## X-squared = 7.1519, df = 2, p-value = 0.02799Again, the low p-value of the Jarque-Bera test indicates that the skewness and kurtosis of the test are not significantly different from their expected values for normal distribution. Thus, the distribution of the model residuals seems statistically not different from a normal distribution.
Antidotes
In order to deal with non-normality in the errors, you could consider the following options:
- Use the log of the dependent variable for you regression.
- For time series data use the first difference of the dependent variable, because you might have a stationarity issue.
- Use a different estimator. For example, if your dependet varible can only have positive values, you might be better off with a generalised linear model, which assumes Gamma-distributed errors. Or for count data you might want to use a Poisson estimator etc.
Heteroskedasticity
Heteroskedasticity arises when the variance of an observation’s error depends on the size of the regressors. Although this does not lead to a bias of the estimated coefficients, it can still render test statistics useless, because their validity relies on the assumption of so-called homoskedastic errors, i.e. the assumption that the errors do not correlate with explanatory variables.
Tests
- Eyeball test: One way to check for heteroskedasticity is to look at a plot, where the absolute values of the residuals - or the squared residuals - are plotted against the regressors of the model. If the size of the absolute (or squared) errors appears to vary with the size of a regressor, this migth indicate heteroskedasticity.
# Create plot input
hetsk_data <- cbind(abs(model$residuals), model$model[, -1])
# Scatterplot matrix
plot(hetsk_data)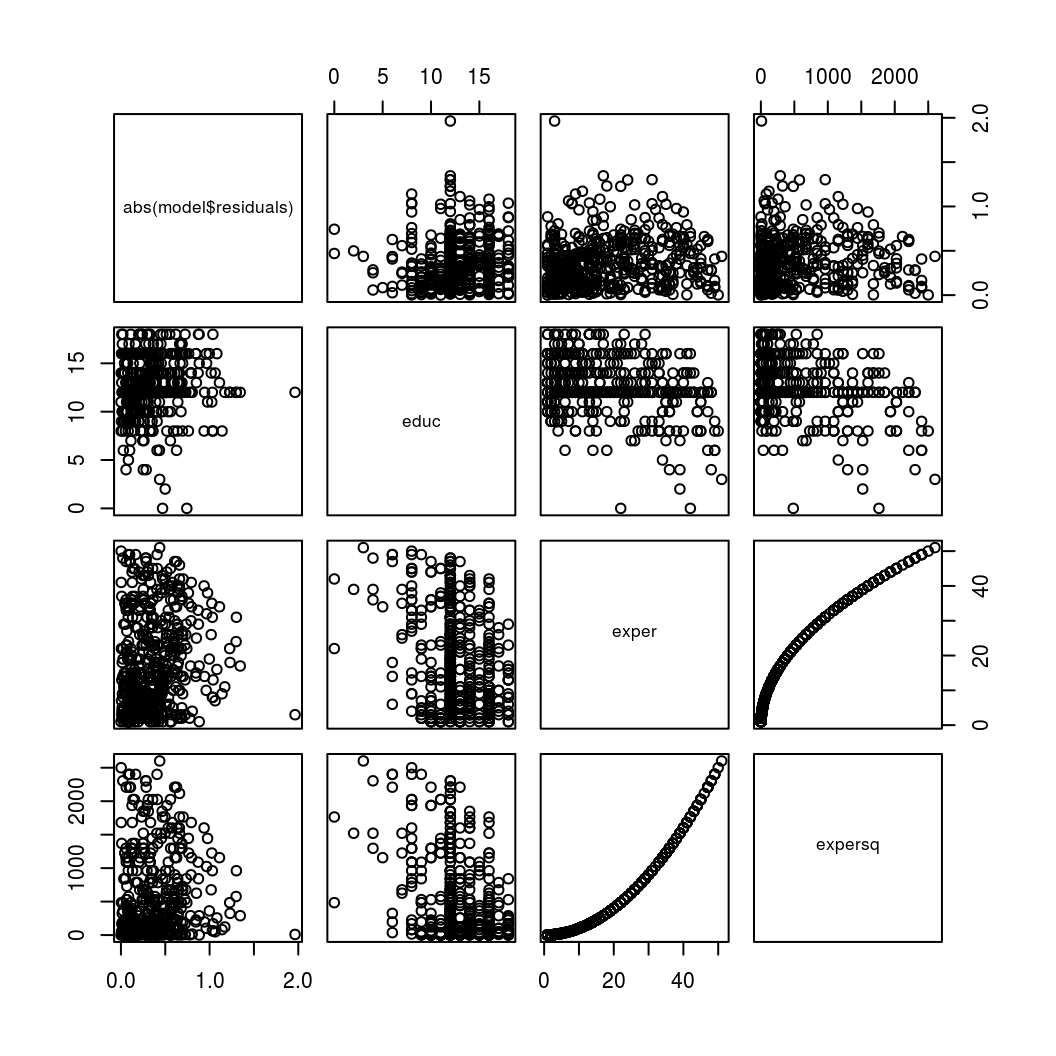
By looking at the first row of the above scatter plot we might notice that the absolute errors seem to be larger for higher values of educ and lower for higher values of expersq. This might warrant to test this more formally.
- The Breusch-Pagan test is implemented in the function
pbtestboth of theAERandlmtestpackage and tests whether the variance of a model’s residuals dependents on the values of the regressors.
library(AER)
bptest(model)##
## studentized Breusch-Pagan test
##
## data: model
## BP = 19.84, df = 3, p-value = 0.0001832- The White test regresses the squared residuals from the original regression model onto a set of regressors that contain the original regressors along with their squares and cross-products. Since it builds on the Breusch-Pagan test, the
bptestcan be used to perform the White test by adding the dependend variables and its squared values to the estimation formula:
bptest(model, ~ lwage + I(lwage^2), data = wage1)##
## studentized Breusch-Pagan test
##
## data: model
## BP = 319.9, df = 2, p-value < 2.2e-16- The Goldfeld-Quandt test basically devides the sample into two groups with low and high values of an explanatory variable and compares the variances of two submodels. If the variances differ, this indicates heteroskedasticity. It is implemented in the function
gqtestof thelmtestpackage:
library(lmtest)
gqtest(model)##
## Goldfeld-Quandt test
##
## data: model
## GQ = 0.74514, df1 = 259, df2 = 259, p-value = 0.9909
## alternative hypothesis: variance increases from segment 1 to 2Note that this test has a series of drawbacks, which is why it is not used very often. The Breusch-Pagan and White test are used much more frequently.
Antidotes
- If the basic structure of heteroskedasticity is unknown - as is usually the case - use heteroskedasticity robust standard errors.
- If the researcher feels that the structure of heteroskedasticity can be estimated, use the feasible GLS estimator
Correlation between explanatory variables and the error term
If explanatory variables are correlated with the error term, i.e. when they are endogenous, an OLS estimator will produce biased estimates. Even if only one variable is causing this problem, the coefficients of all other variables will be affected as well. Since economic data is frequently affected by this problem - due to, for example, measurement error in explanatory variables, autoregression with autocorrelated errors, simultaneity, omitted explanatory variables or sample selection - it is important to test for it.
Hausman test
A popular method to test for correlation between explanatory variables and the error term is the Hausman test. It basically tests whether the results of the IV estimator are significantly different from the OLS estimator. To illustrate this, let’s use the wage2 dataset from the wooldridge package and, again, estimate a basic Mincer earnings function as described above. However, this time we add IQ as a measure of ability. But since it is assumed that the measure is correlated with the error term, we use an additional measure of ability, KWW, as an instrument for IQ.
library(dplyr)
library(wooldridge)
data("wage2")
# Add squared experience "expersq"
wage2 <- wage2 %>%
mutate(expersq = exper^2)
# IV estimator
iv_model <- ivreg(lwage ~ educ + exper + expersq + IQ | KWW + educ + exper + expersq,
data = wage2)The summary function of the ivreg object, performs the Hausman test automatically. We only have to add the argument diagnostics = TRUE.
summary(iv_model, diagnostics = TRUE)##
## Call:
## ivreg(formula = lwage ~ educ + exper + expersq + IQ | KWW + educ +
## exper + expersq, data = wage2)
##
## Residuals:
## Min 1Q Median 3Q Max
## -2.097500 -0.256972 0.005075 0.274838 1.284012
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 4.5137035 0.2437555 18.517 < 2e-16 ***
## educ 0.0096496 0.0155761 0.620 0.536
## exper 0.0144617 0.0145717 0.992 0.321
## expersq 0.0001935 0.0006100 0.317 0.751
## IQ 0.0191398 0.0038791 4.934 9.54e-07 ***
##
## Diagnostic tests:
## df1 df2 statistic p-value
## Weak instruments 1 930 77.13 < 2e-16 ***
## Wu-Hausman 1 929 15.63 8.26e-05 ***
## Sargan 0 NA NA NA
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 0.4231 on 930 degrees of freedom
## Multiple R-Squared: -0.004854, Adjusted R-squared: -0.009176
## Wald test: 36.38 on 4 and 930 DF, p-value: < 2.2e-16The (Wu-)Hausman test statistic has a very low p-value and, hence, the test rejects the null of no correcation between IQ and the error term in this example.
Sargan test
The Sargan test checks whether an instrument is correlated with the errors, which should not be the case for good instruments. In the example above, the test statistic of the Sargan test is NA. This is to be expected, because we use one instrument for one endogenous regressor and the mathematical properties of the test require us to have at least one instrument more than endogenous regressor. So, if we use more instruments than endogenous regressors, the system would be identified and we would be able to obtain a test statistic. A low statistic (with rather high p-value) would indicate that the instrument is uncorrelated with the error term.
To be continued… (January 2020)
Literature
Kennedy, P. (2014). A Guide to Econometrics. Malden (Mass.): Blackwell Publishing 6th ed.
Wooldridge, J.M. (2013). Introductory econometrics: A modern approach. (5thed.)